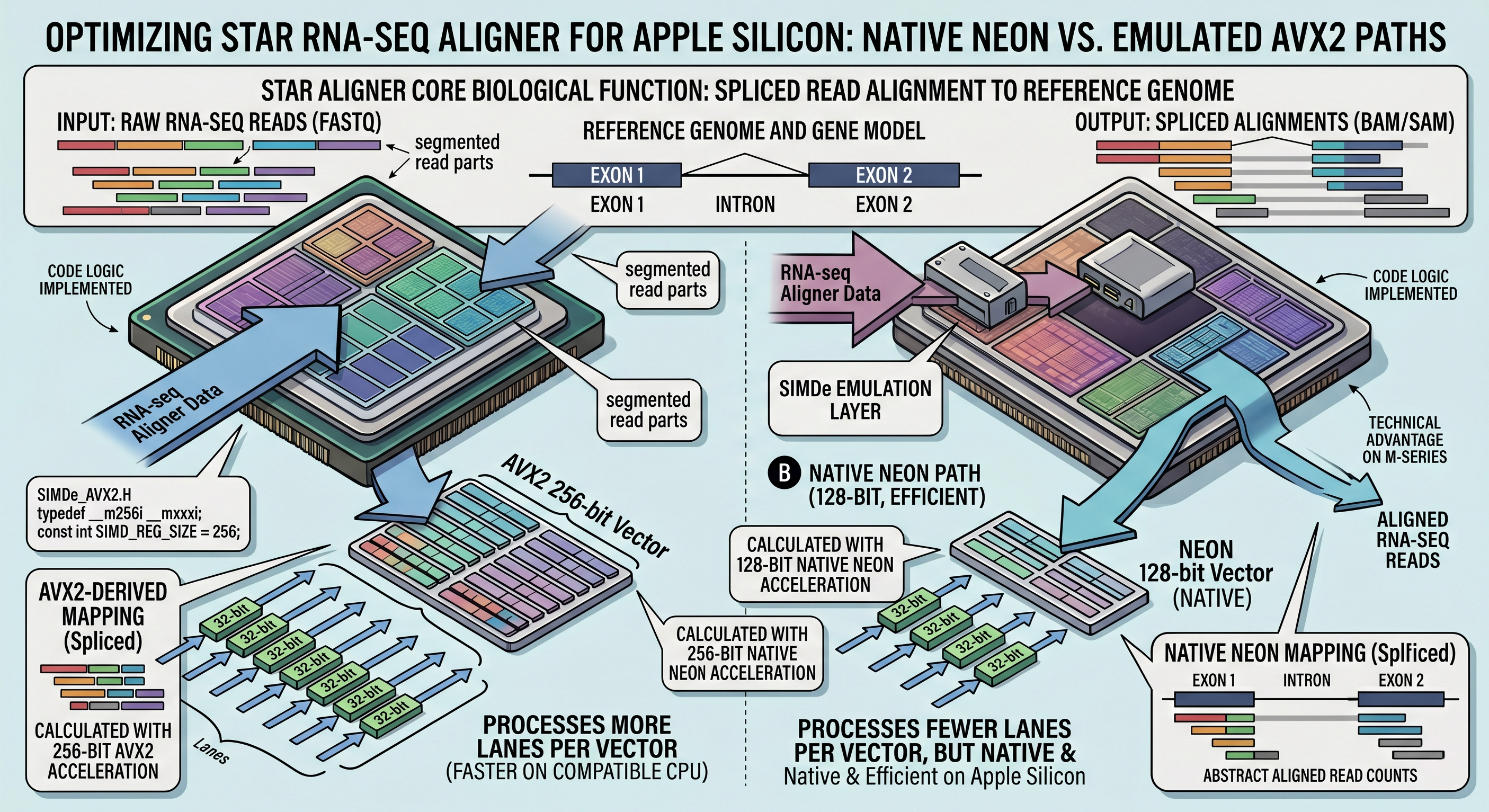
Added NEON support to STAR RNA-seq aligner tool.
Bioinformatics
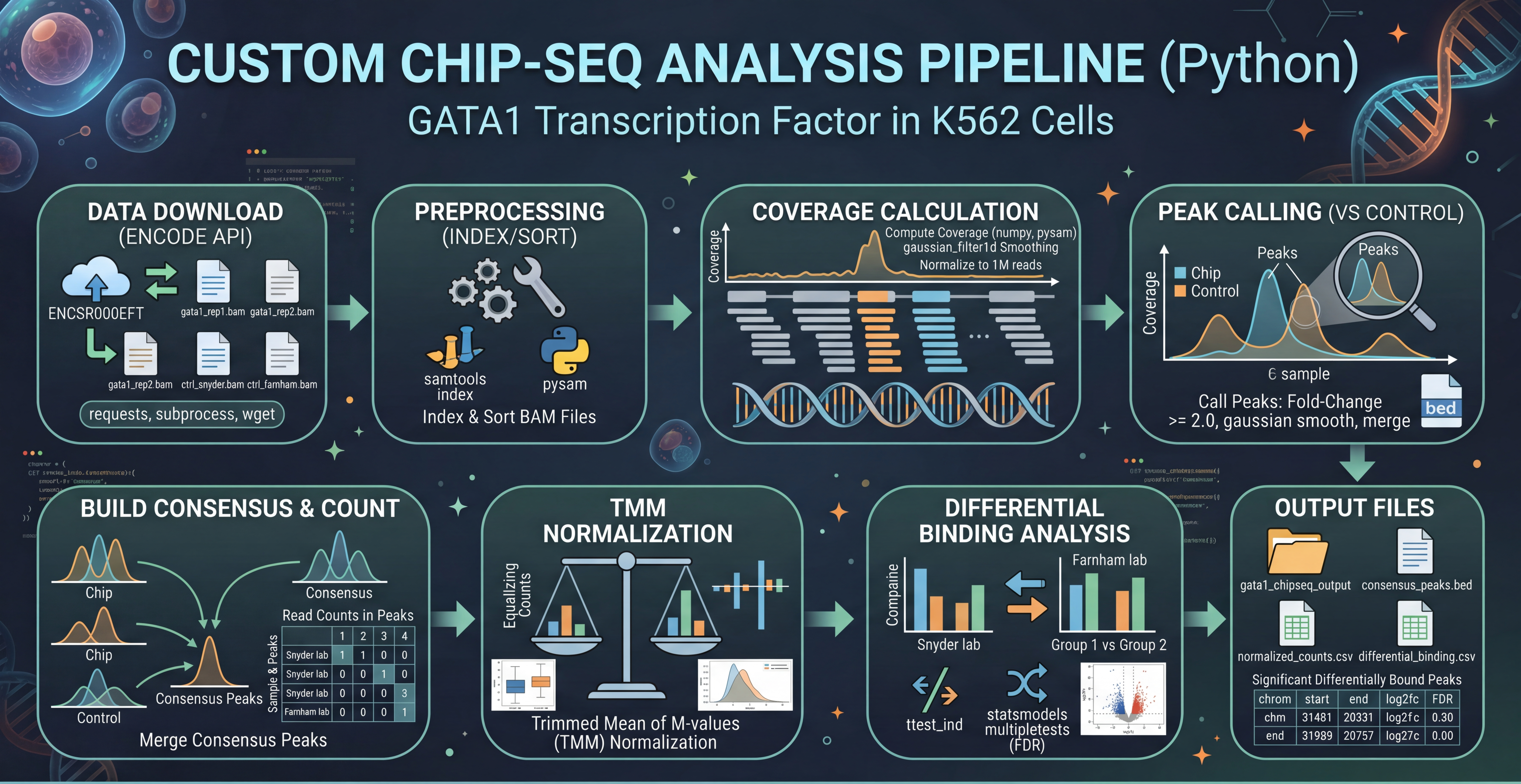
Downloading and analyzing ChIP-seq data.
Bioinformatics
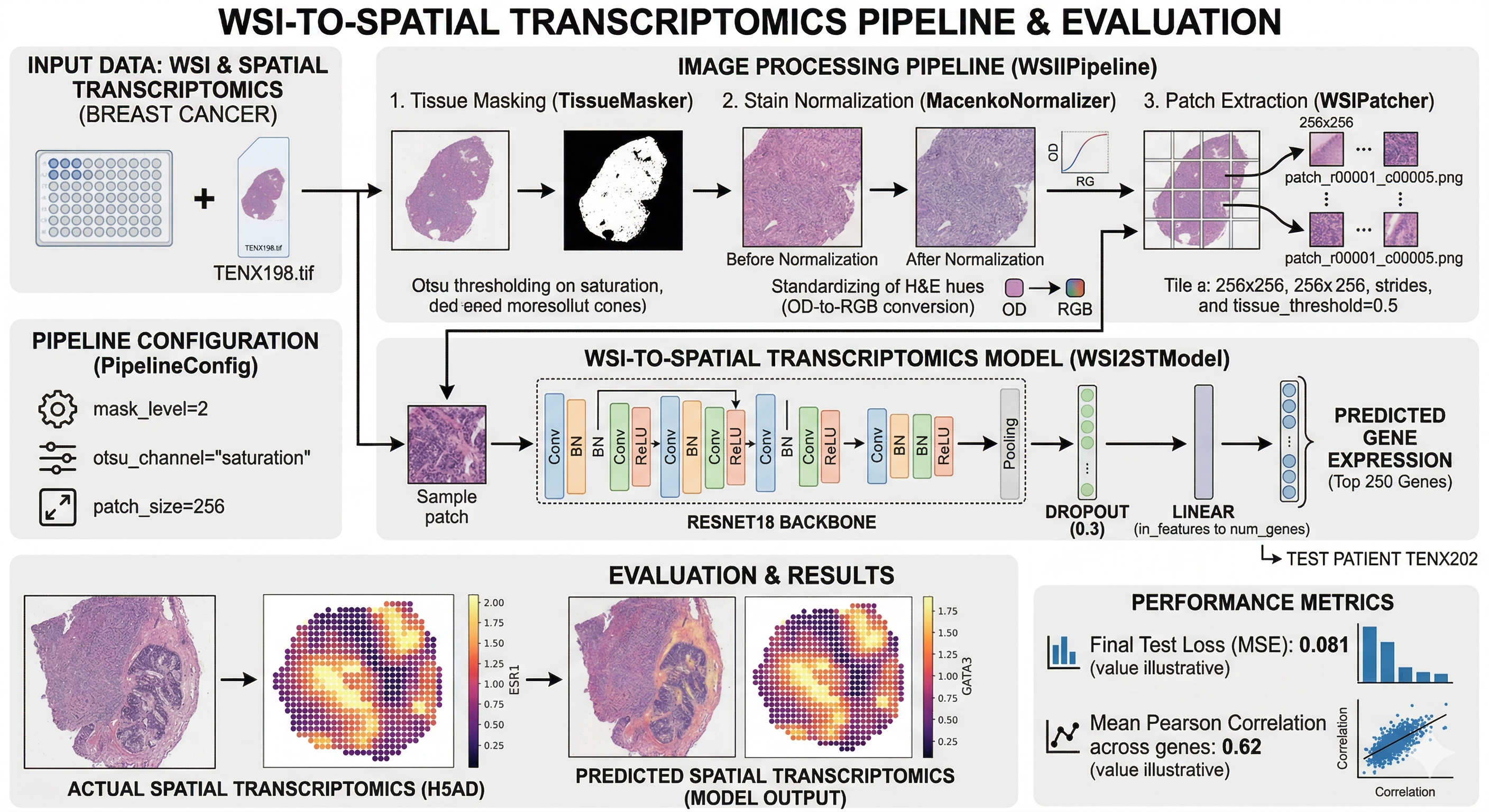
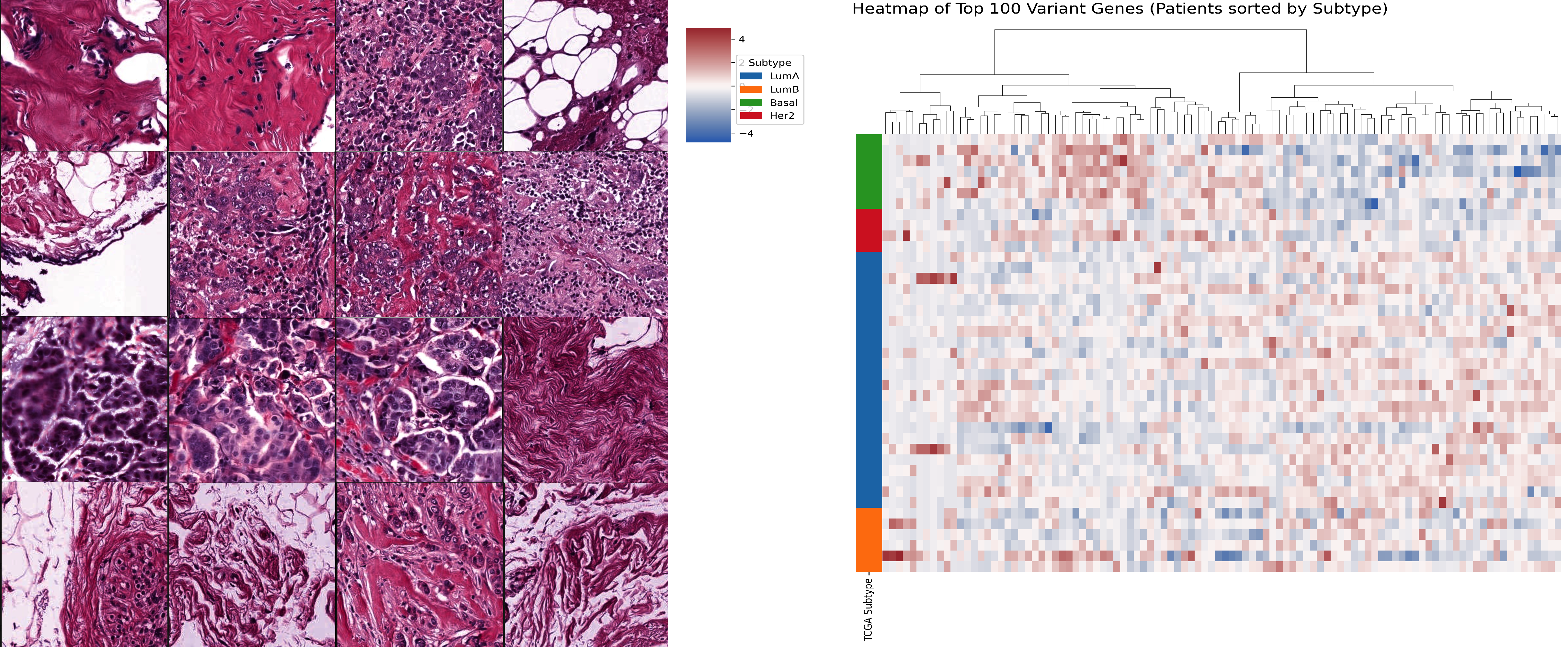
Predicting Spatial Transcriptomics (ST) using Whole Slide Images (WSI).
Bioinformatics
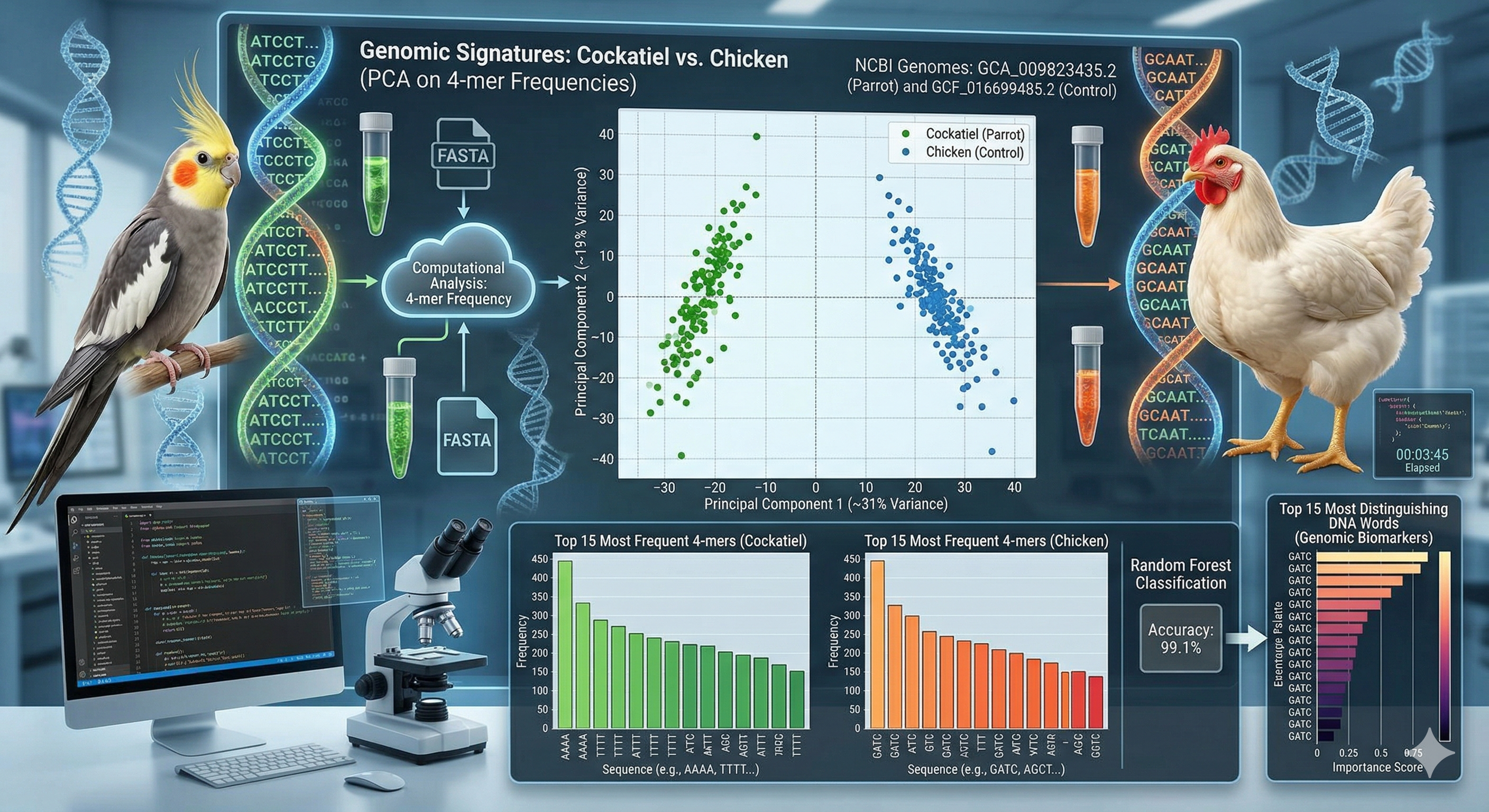
Genetic differences between the Cockatiel and a biological control species, the Chicken.
Bioinformatics
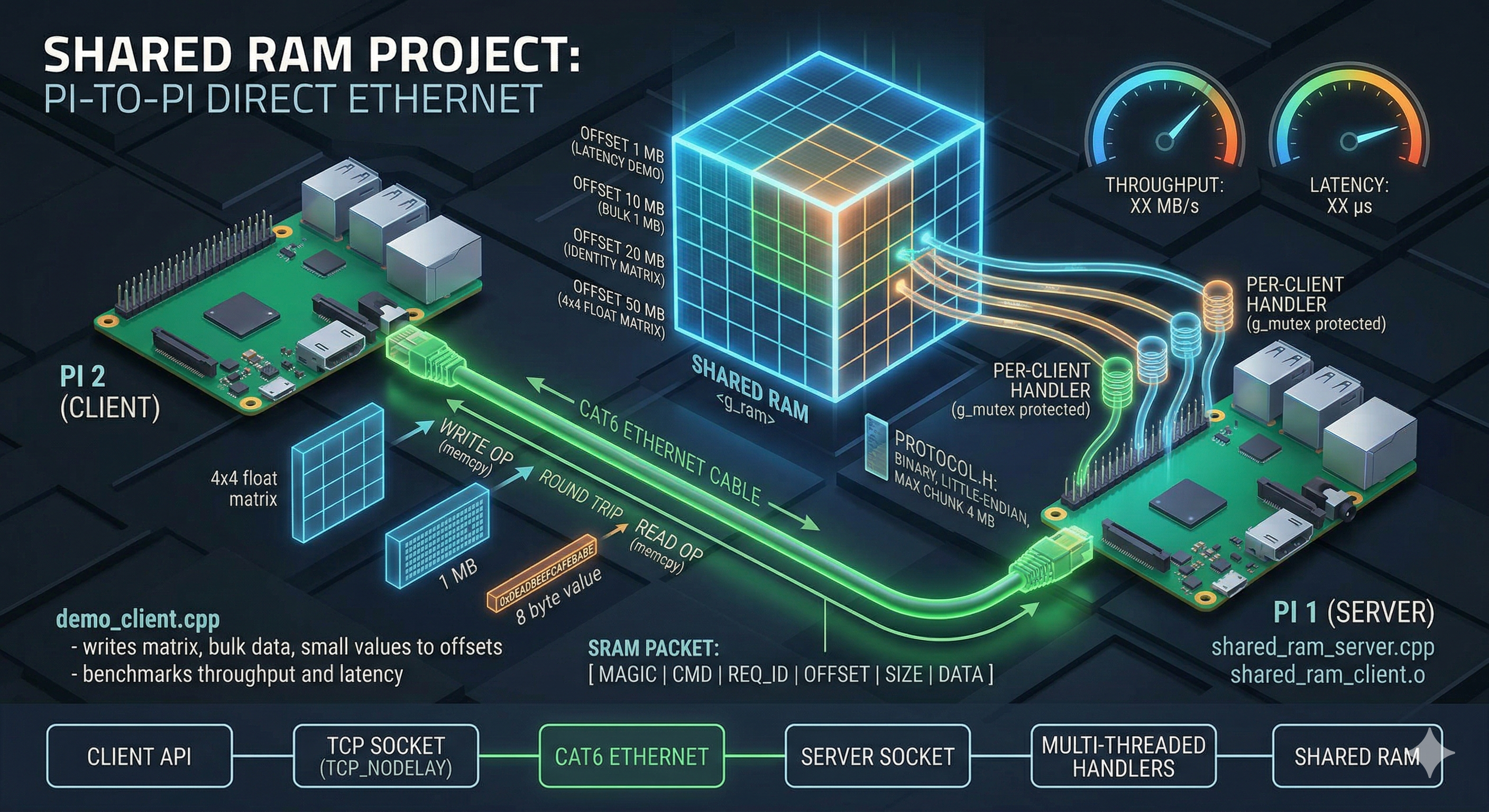
A Custom Message Passing Interface (MPI) with Remote Direct Memory Access (RDMA).
MPIRDMA
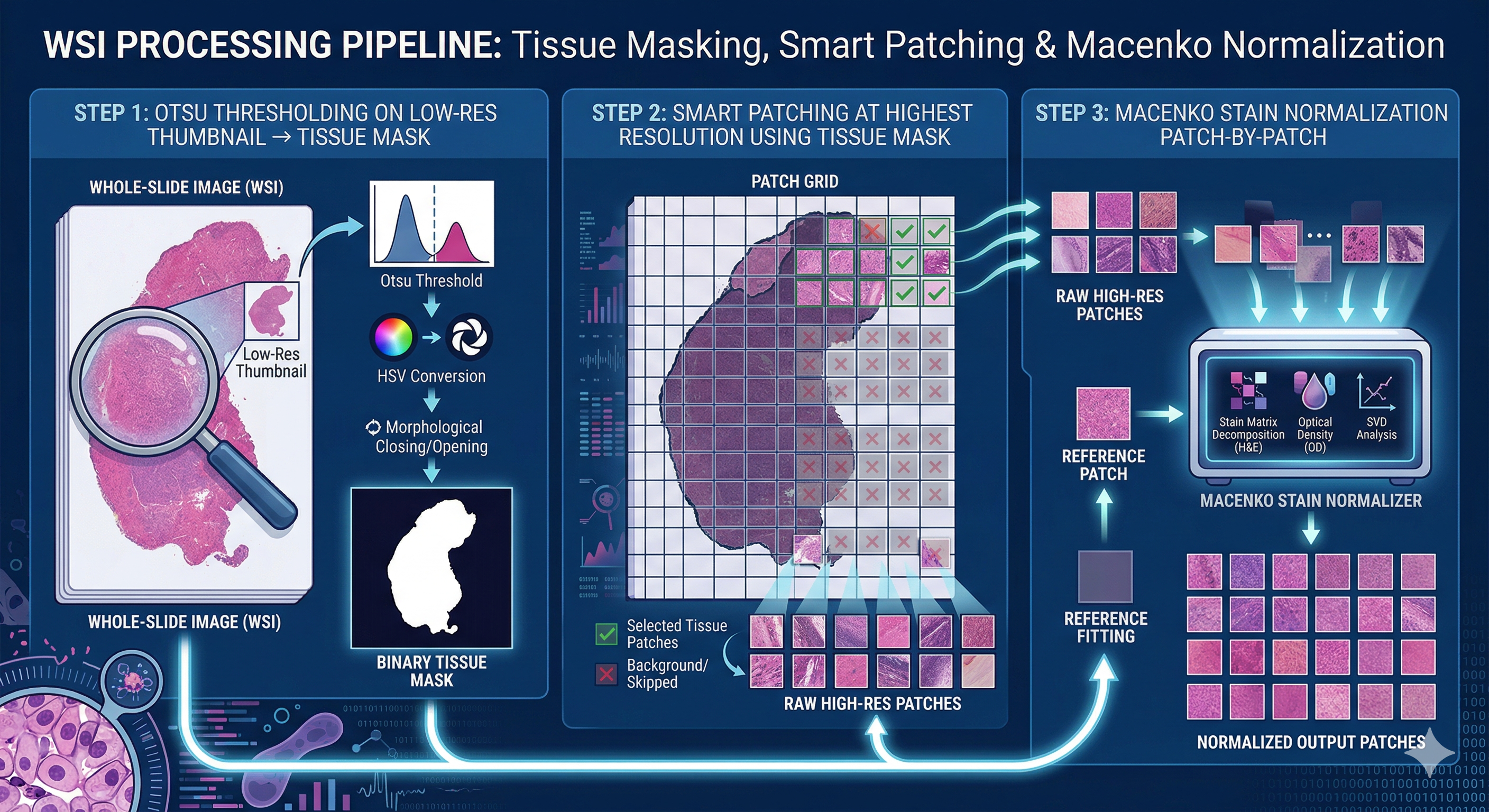
WSI Data Preprocessing Pipeline.
Histopathology
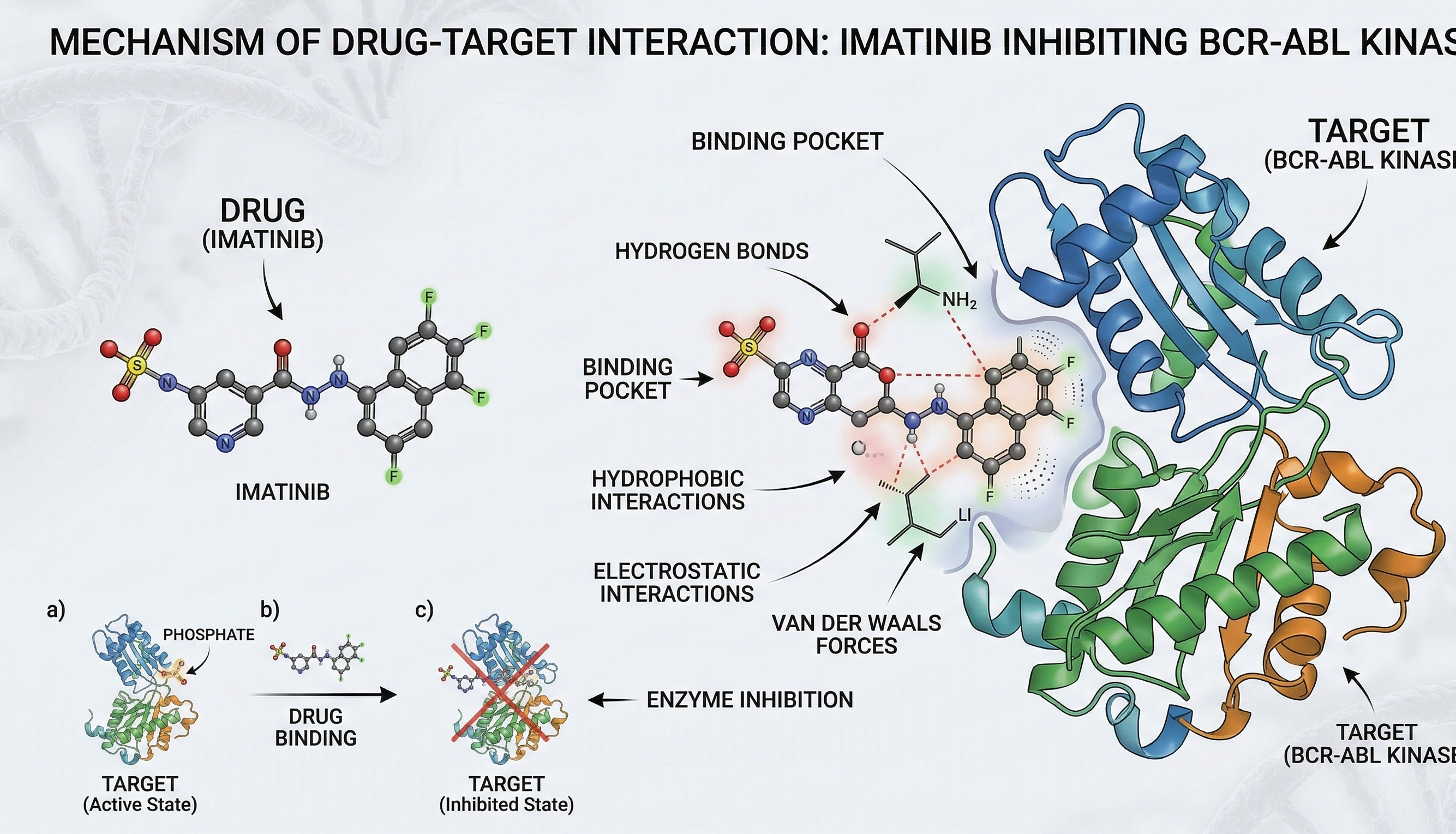
Affinity (pKd) regression using Deep Neural Networks.
BioinformaticsDrug-Target Interaction
A multi-agent, multimodal RAG, AI dermatological assistant.
BioinformaticsRAG
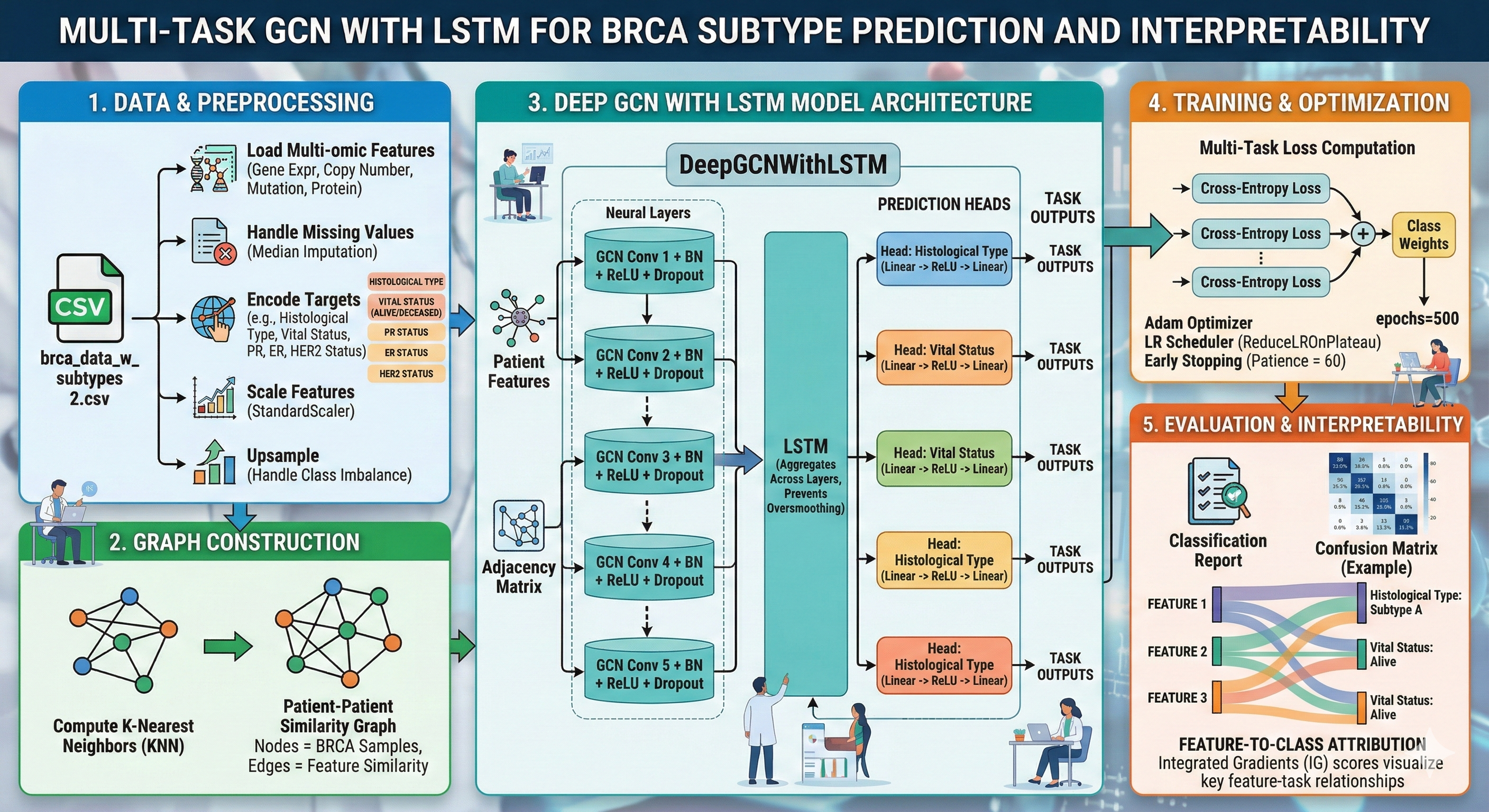
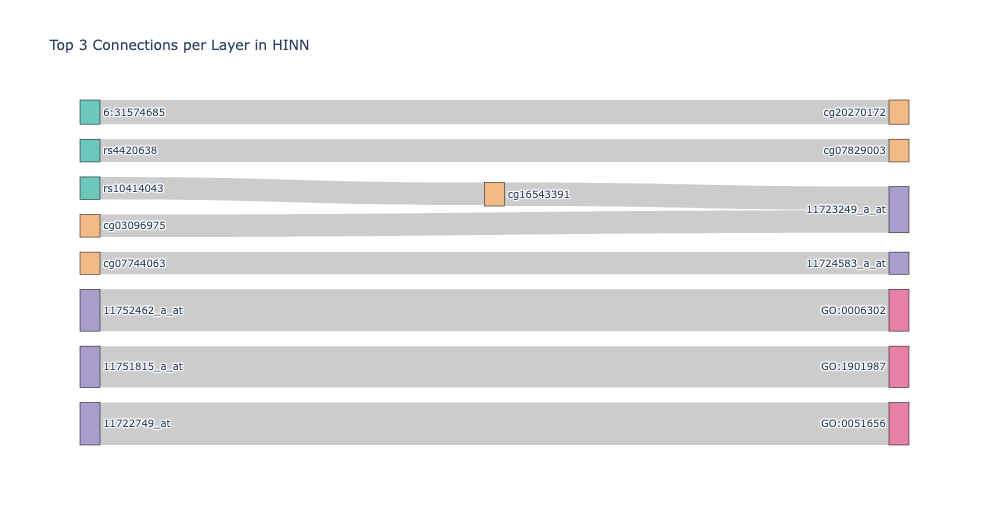
A model to predict histological, vital, and receptor status by concatenating 1,936 multi-omics features per patient node.
BioinformaticsPyTorch
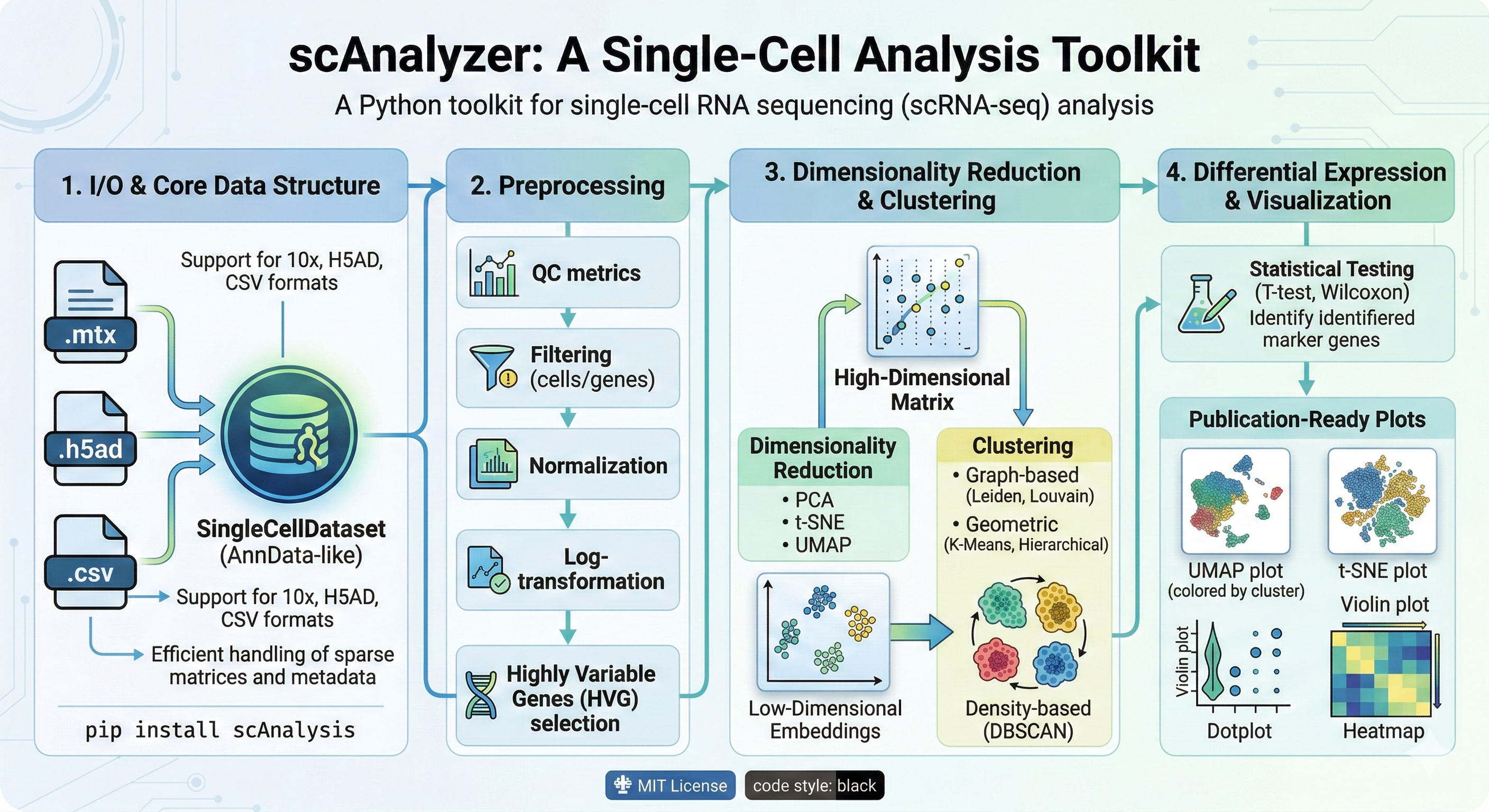
A Python toolkit for single-cell RNA sequencing (scRNA-seq) analysis.
Bioinformatics
FASTQ k-merization
Bioinformatics
RNA sequence data normalization.
RBioinformatics
Exon Junction Analyzer.
PyTorchBioinformaticsSTAR
Single cell RNA sequencing using GenePT.
PyTorchBioinformatics
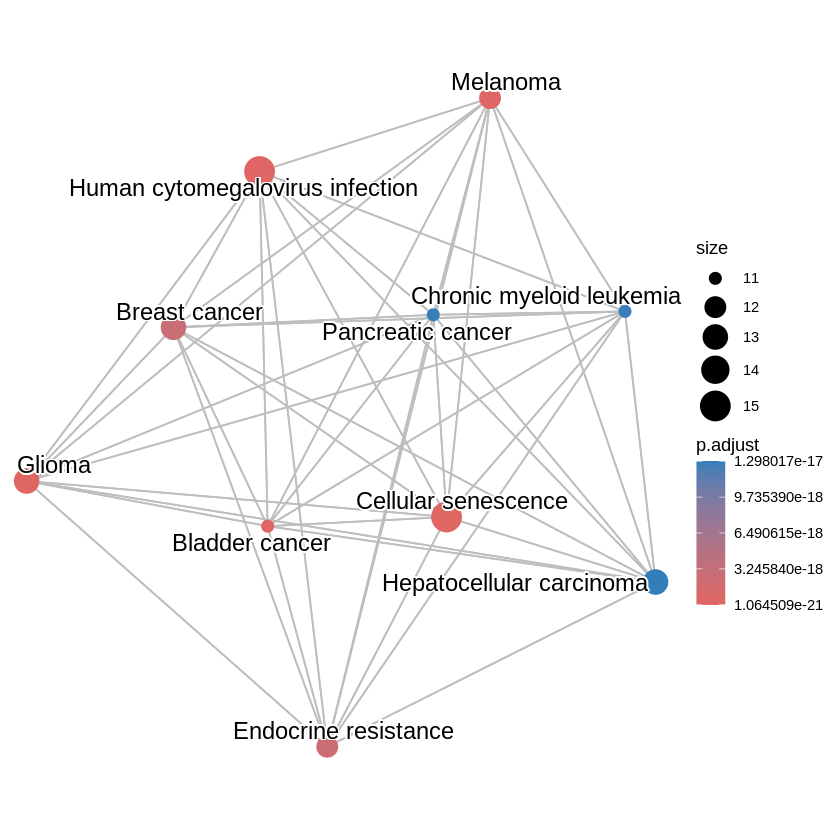
KEGG Pathway Enrichment Analysis using R and clusterProfiler.
RBioinformatics
Histopathologic CapsuleNet for cancer classification.
PyTorchBioinformatics
This Python script implements a genetic algorithm that recreates a target image (the Mona Lisa) by iteratively evolving a population of 500 semi-transparent circles.
PyTorchBioinformatics
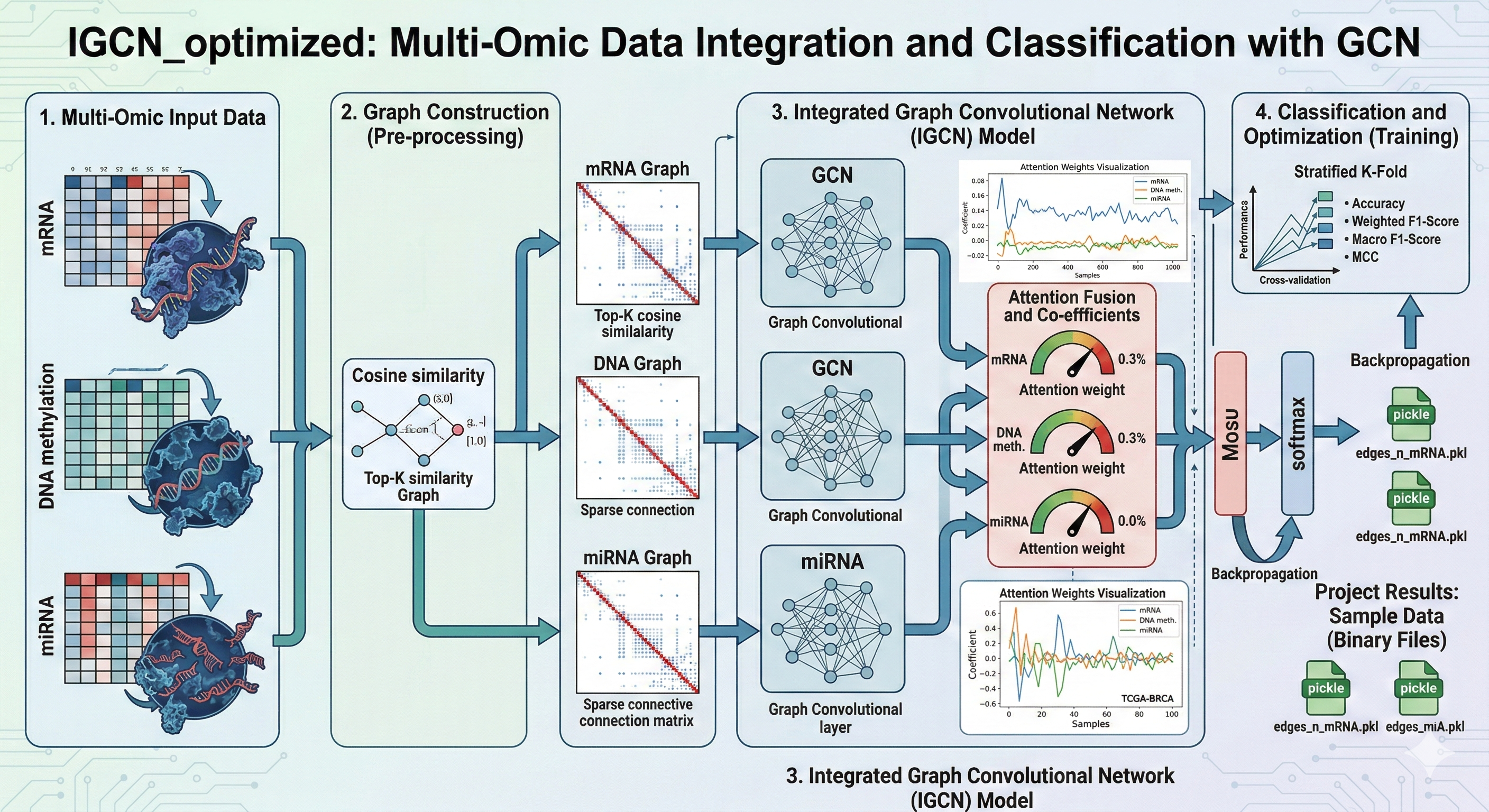
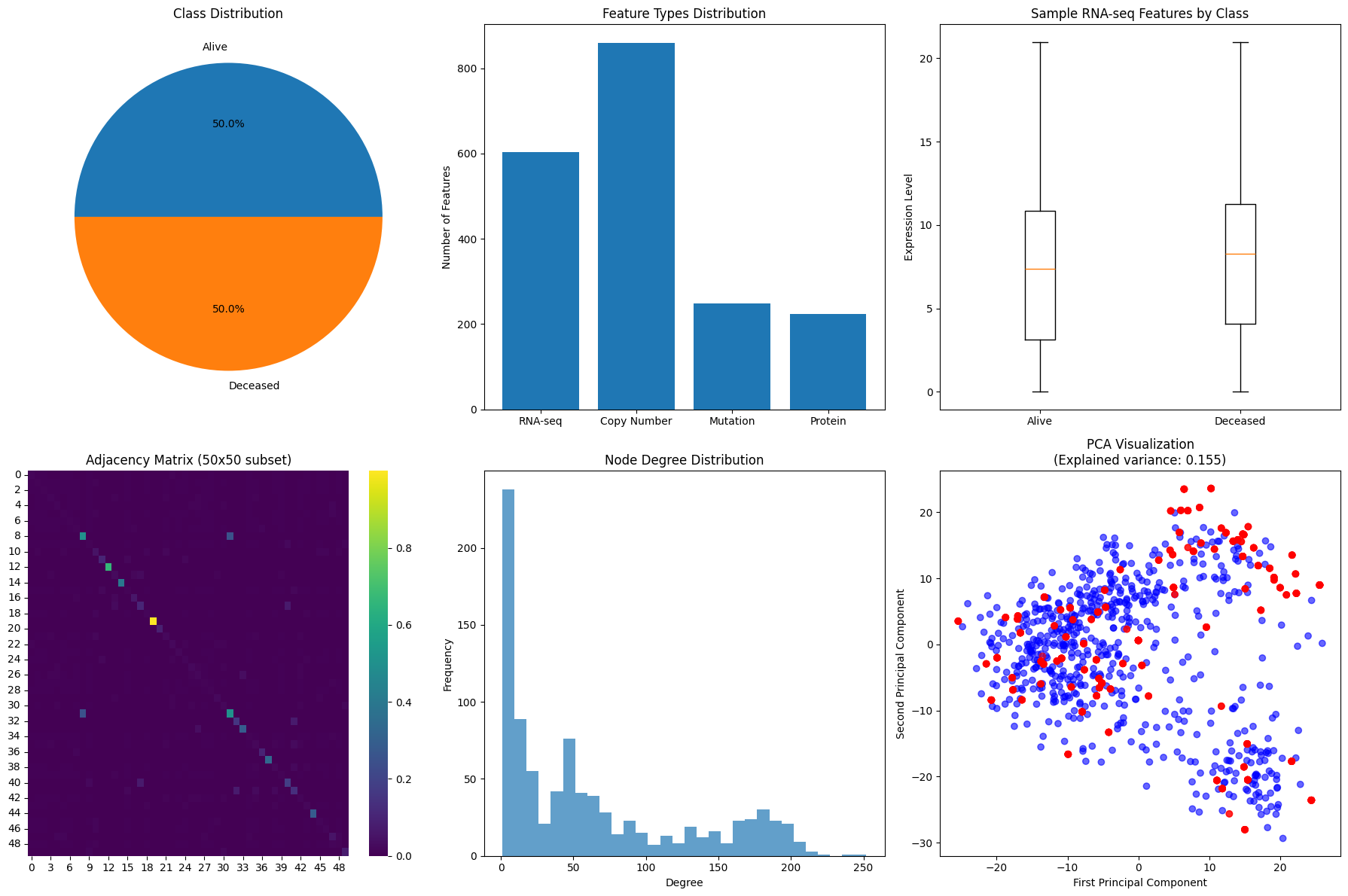
This Jupyter Notebook implements a graph neural network (GNN) pipeline for binary survival classification using multi-omics breast cancer data.
TensorFlowMedical AI
Design novel protein sequences with unique binding sites using GPT-2.
PyTorchMedical AITransformers
DINOv2 Nuclei Segmentation.
PythonMedical AI
A Gradio app for Microsoft's SOTA grounded radiology report generation model.
VisualizationMedical AI
Fine tuned SmolLM-135M-Instruct using Group Relative Policy Optimization.
LLMsGRPO
Fine tuned Llama 2 on a custom product review sentiment dataset with instructions.
LLMsInstruction Tuning
A Gradio app for TotalSegmentator.
VisualizationMedical AI
Fine-tuned the model.
Medical AIPython
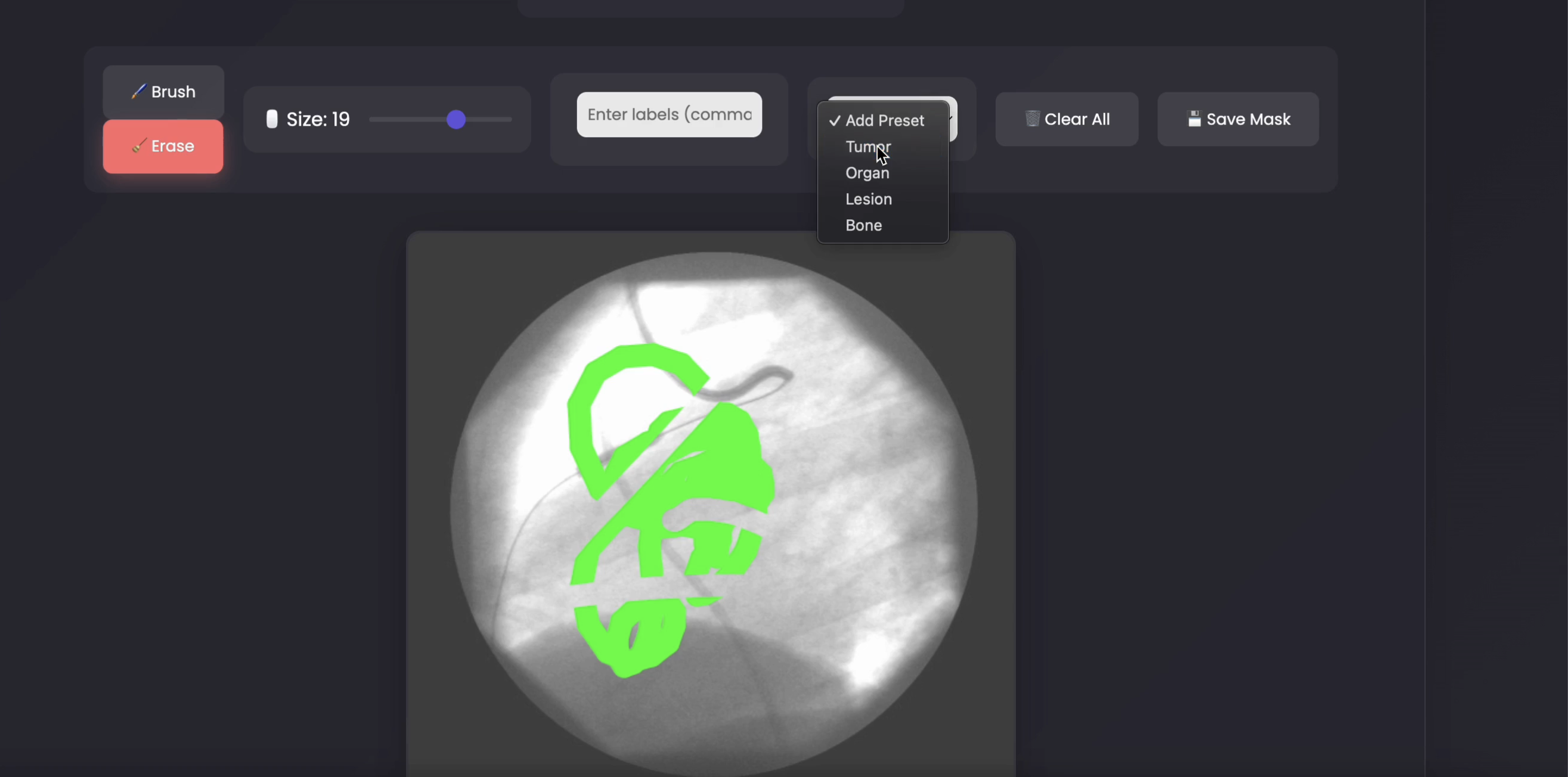
A DICOM annotation tool for 3D segmentation.
Medical AIVisualization
Speech emotion classification with multi-head self-attention.
Audio classification
A Gradio app for SigLIP 2 Zero-shot image classification.
Python