import py3Dmol
import glob
import matplotlib.pyplot as plt
from colabfold.colabfold import plot_plddt_legend
from colabfold.colabfold import pymol_color_list, alphabet_list
rank_num = 1 #["1", "2", "3", "4", "5"]
color = "lDDT" # ["chain", "lDDT", "rainbow"]
show_sidechains = True
show_mainchains = True
tag = results["rank"][0][rank_num - 1]
jobname_prefix = ".custom" if msa_mode == "custom" else ""
pdb_filename = f"{jobname}/{jobname}{jobname_prefix}_unrelaxed_{tag}.pdb"
pdb_file = glob.glob(pdb_filename)
def show_pdb(rank_num=1, show_sidechains=False, show_mainchains=False, color="lDDT"):
model_name = f"rank_{rank_num}"
view = py3Dmol.view(js='https://3dmol.org/build/3Dmol.js',)
view.addModel(open(pdb_file[0],'r').read(),'pdb')
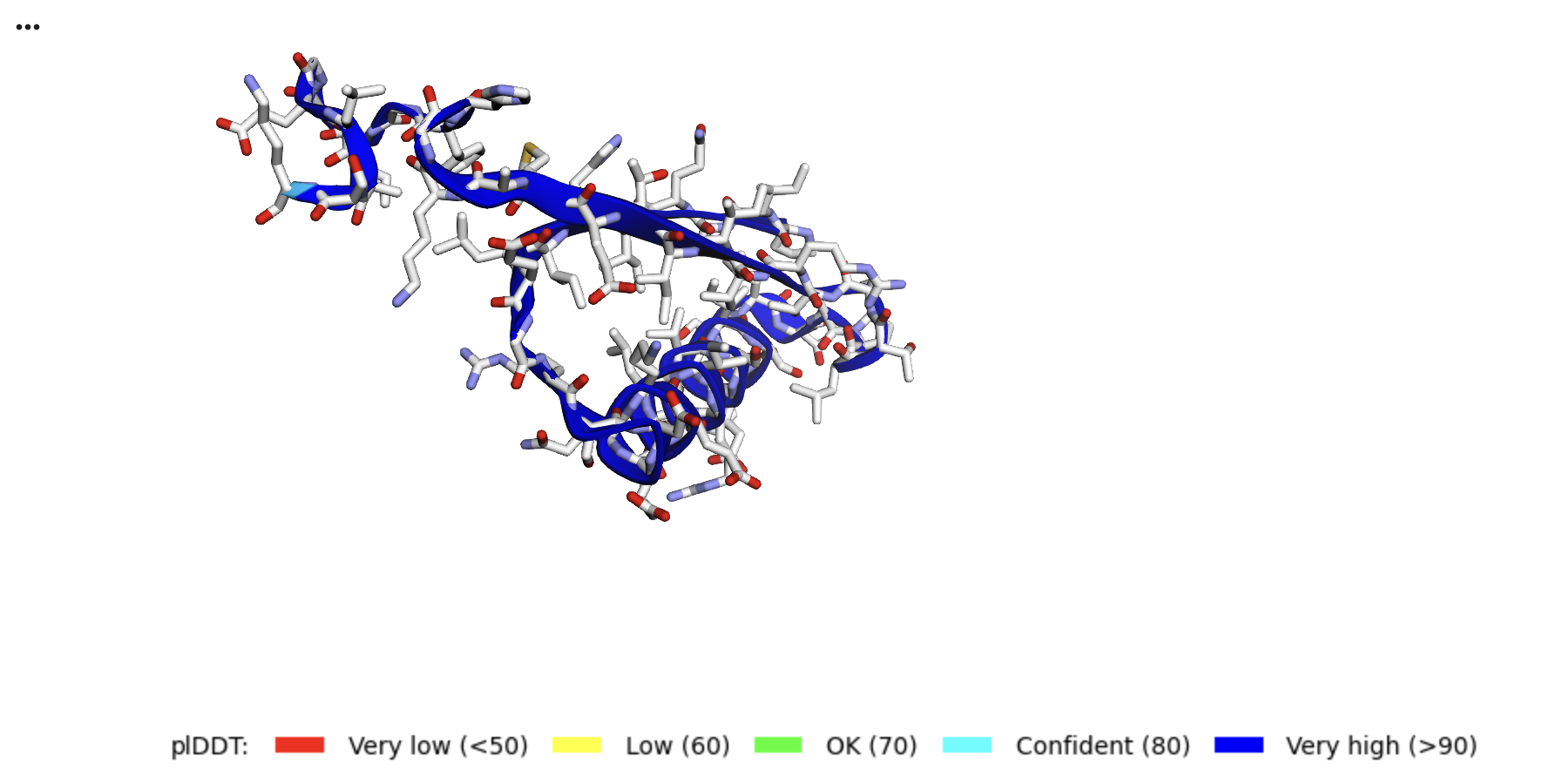
if color == "lDDT":
view.setStyle({'cartoon': {'colorscheme': {'prop':'b','gradient': 'roygb','min':50,'max':90}}})
elif color == "rainbow":
view.setStyle({'cartoon': {'color':'spectrum'}})
elif color == "chain":
chains = len(queries[0][1]) + 1 if is_complex else 1
for n,chain,color in zip(range(chains),alphabet_list,pymol_color_list):
view.setStyle({'chain':chain},{'cartoon': {'color':color}})
if show_sidechains:
BB = ['C','O','N']
view.addStyle({'and':[{'resn':["GLY","PRO"],'invert':True},{'atom':BB,'invert':True}]},
{'stick':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
view.addStyle({'and':[{'resn':"GLY"},{'atom':'CA'}]},
{'sphere':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
view.addStyle({'and':[{'resn':"PRO"},{'atom':['C','O'],'invert':True}]},
{'stick':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
if show_mainchains:
BB = ['C','O','N','CA']
view.addStyle({'atom':BB},{'stick':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
view.zoomTo()
return view
show_pdb(rank_num, show_sidechains, show_mainchains, color).show()
if color == "lDDT":
plot_plddt_legend().show()